Model
Architecture
X-Cell is a set-level diffusion transformer that predicts perturbed gene expression profiles from control cell populations. Unlike autoregressive single-cell foundation models, X-Cell operates on sets of cells and is trained explicitly on interventional data via distribution-matching objectives.
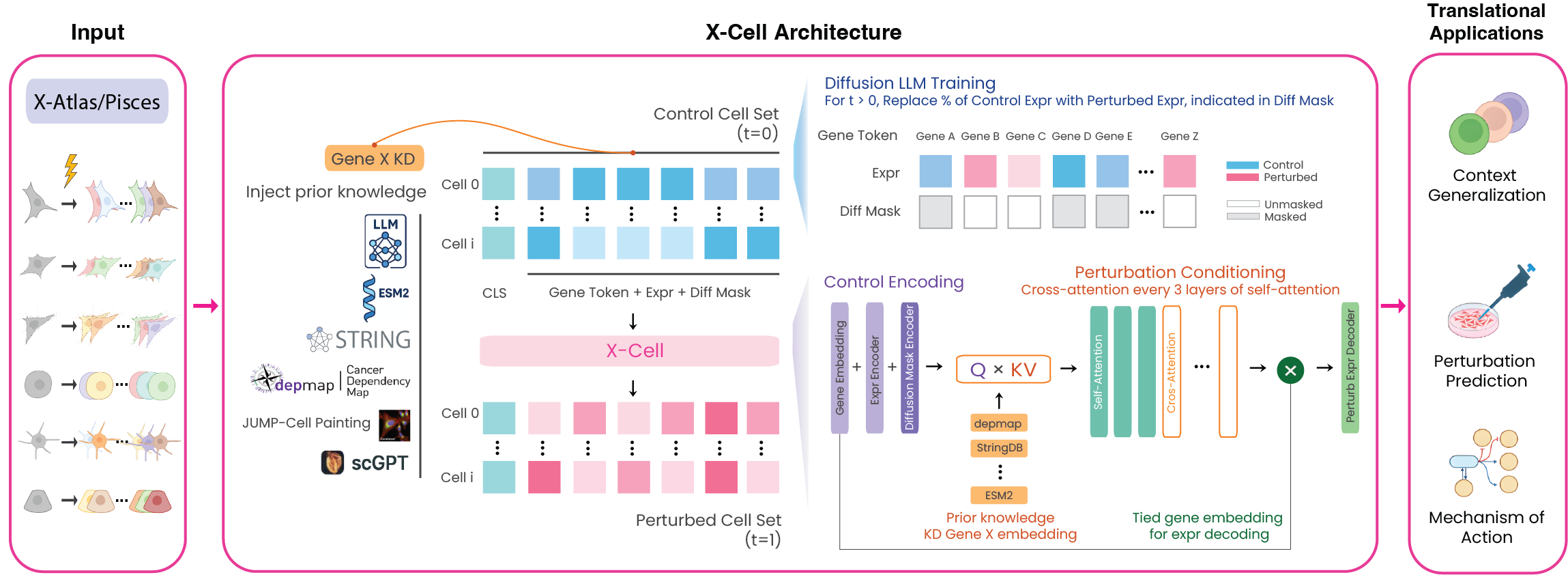
Key design choices
Diffusion-based training. Each training sample has a random fraction (25%, 50%, 75%, or 100%) of control gene expression positions replaced with ground-truth perturbed values. The model learns to predict the full perturbed profile from this partially revealed input. At inference, predictions are iteratively refined across 4 steps (coarse-to-fine).
Multi-modal biological priors via cross-attention. At every third self-attention layer, Flamingo-style cross-attention conditions gene representations on six prior knowledge tokens per perturbation:
| Source | Content | Dimension |
|---|---|---|
| ESM-2 | Protein language model embeddings | 5120 |
| STRING | Protein-protein interaction network | 512 |
| GenePT | LLM gene representations | 3072 |
| DepMap | Genetic dependency profiles | 1150 |
| JUMP-Cell Painting | Morphological features | 259 |
| Gene identity | Stop-gradient gene embedding | — |
Tied output embeddings. The output head projects back through the shared gene embedding matrix (PaLM-style 1/√d scaling), acting as an implicit regularizer against conservative collapse.
X-Cell Mini
| X-Cell Mini | |
|---|---|
| Parameters | 55M |
| Layers | 12 |
| Hidden dim | 512 |
| Attention heads | 8 |
| FFN | ReLU, 1× |
| Normalization | Post-LN (LayerNorm) |
| Cross-attn layers | 4 |
| Init | scGPT |
| Training | Replogle-Nadig |
| Min GPU VRAM | 8 GB (1 GPU) |
Scaling
X-Cell follows power-law scaling consistent with large language models. Train loss scales as L(N) ∝ N⁻⁰·³² (α = 0.32, R² = 0.96) across five model sizes from 83M to 3.1B parameters.
Weights
Model weights are hosted on HuggingFace:
 Open on HuggingFace
Open on HuggingFace